Classification
[1]:
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout
from tensorflow.keras.optimizers import SGD
from freeforestml import Variable, Process, Cut, \
HepNet, ClassicalCV, EstimatorNormalizer, \
HistogramFactory, confusion_matrix, atlasify, \
McStack
from freeforestml import toydata, example_style
example_style()
2023-08-02 16:28:32.774772: I tensorflow/core/platform/cpu_feature_guard.cc:193] This TensorFlow binary is optimized with oneAPI Deep Neural Network Library (oneDNN) to use the following CPU instructions in performance-critical operations: AVX2 AVX512F FMA
To enable them in other operations, rebuild TensorFlow with the appropriate compiler flags.
2023-08-02 16:28:32.900177: W tensorflow/compiler/xla/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'libcudart.so.11.0'; dlerror: libcudart.so.11.0: cannot open shared object file: No such file or directory
2023-08-02 16:28:32.900207: I tensorflow/compiler/xla/stream_executor/cuda/cudart_stub.cc:29] Ignore above cudart dlerror if you do not have a GPU set up on your machine.
2023-08-02 16:28:33.621320: W tensorflow/compiler/xla/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'libnvinfer.so.7'; dlerror: libnvinfer.so.7: cannot open shared object file: No such file or directory
2023-08-02 16:28:33.621424: W tensorflow/compiler/xla/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'libnvinfer_plugin.so.7'; dlerror: libnvinfer_plugin.so.7: cannot open shared object file: No such file or directory
2023-08-02 16:28:33.621436: W tensorflow/compiler/tf2tensorrt/utils/py_utils.cc:38] TF-TRT Warning: Cannot dlopen some TensorRT libraries. If you would like to use Nvidia GPU with TensorRT, please make sure the missing libraries mentioned above are installed properly.
[2]:
df = toydata.get()
[3]:
p_ztt = Process(r"$Z\rightarrow\tau\tau$", range=(0, 0))
p_sig = Process(r"Signal", range=(1, 1))
s_all = McStack(p_ztt, p_sig)
[4]:
hist_factory = HistogramFactory(df, stacks=[s_all], weight="weight")
Cut-based
First, we set up a cut-based event selection as a benchmark.
[5]:
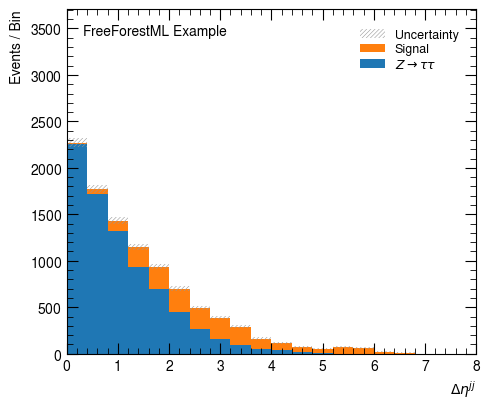
hist_factory(Variable("$\Delta \eta^{jj}$",
lambda d: (d.jet_1_eta - d.jet_2_eta).abs()),
bins=20, range=(0, 8))
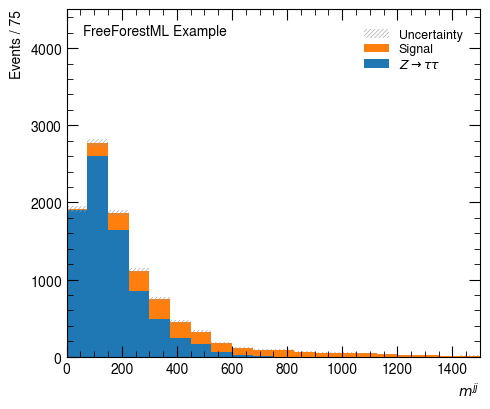
hist_factory(Variable("$m^{jj}$", "m_jj"),
bins=20, range=(0, 1500))
None
[6]:
c_sr = Cut(lambda d: d.m_jj > 400) & \
Cut(lambda d: d.jet_2_pt >= 30) & \
Cut(lambda d: d.jet_1_eta * d.jet_2_eta < 0) & \
Cut(lambda d: (d.jet_2_eta - d.jet_1_eta).abs() > 3)
c_sr.label = "Signal"
c_rest = (~c_sr)
c_rest.label = "Rest"
[7]:
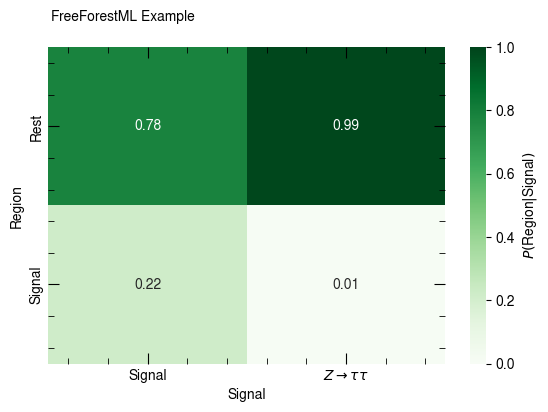
confusion_matrix(df, [p_sig, p_ztt], [c_sr, c_rest],
x_label="Signal", y_label="Region", annot=True, weight="weight")
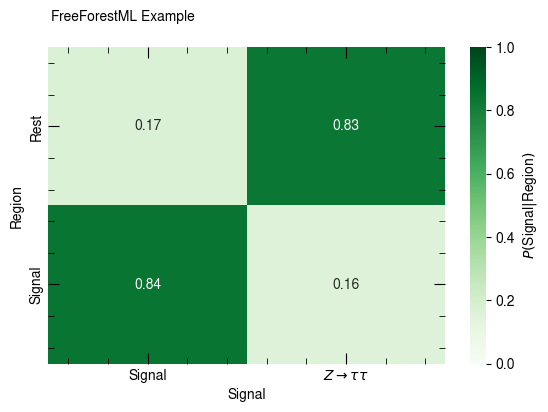
confusion_matrix(df, [p_sig, p_ztt], [c_sr, c_rest], normalize_rows=True,
x_label="Signal", y_label="Region", annot=True, weight="weight")
None
Neural Network
[8]:
df['dijet_deta'] = (df.jet_1_eta - df.jet_2_eta).abs()
df['dijet_prod_eta'] = (df.jet_1_eta * df.jet_2_eta)
input_var = ['dijet_prod_eta', 'm_jj', 'dijet_deta', 'higgs_pt', 'jet_2_pt', 'jet_1_eta', 'jet_2_eta', 'tau_eta']
output_var = ['is_sig', 'is_ztt']
[9]:
df["is_sig"] = p_sig.selection.idx_array(df)
df["is_ztt"] = p_ztt.selection.idx_array(df)
[10]:
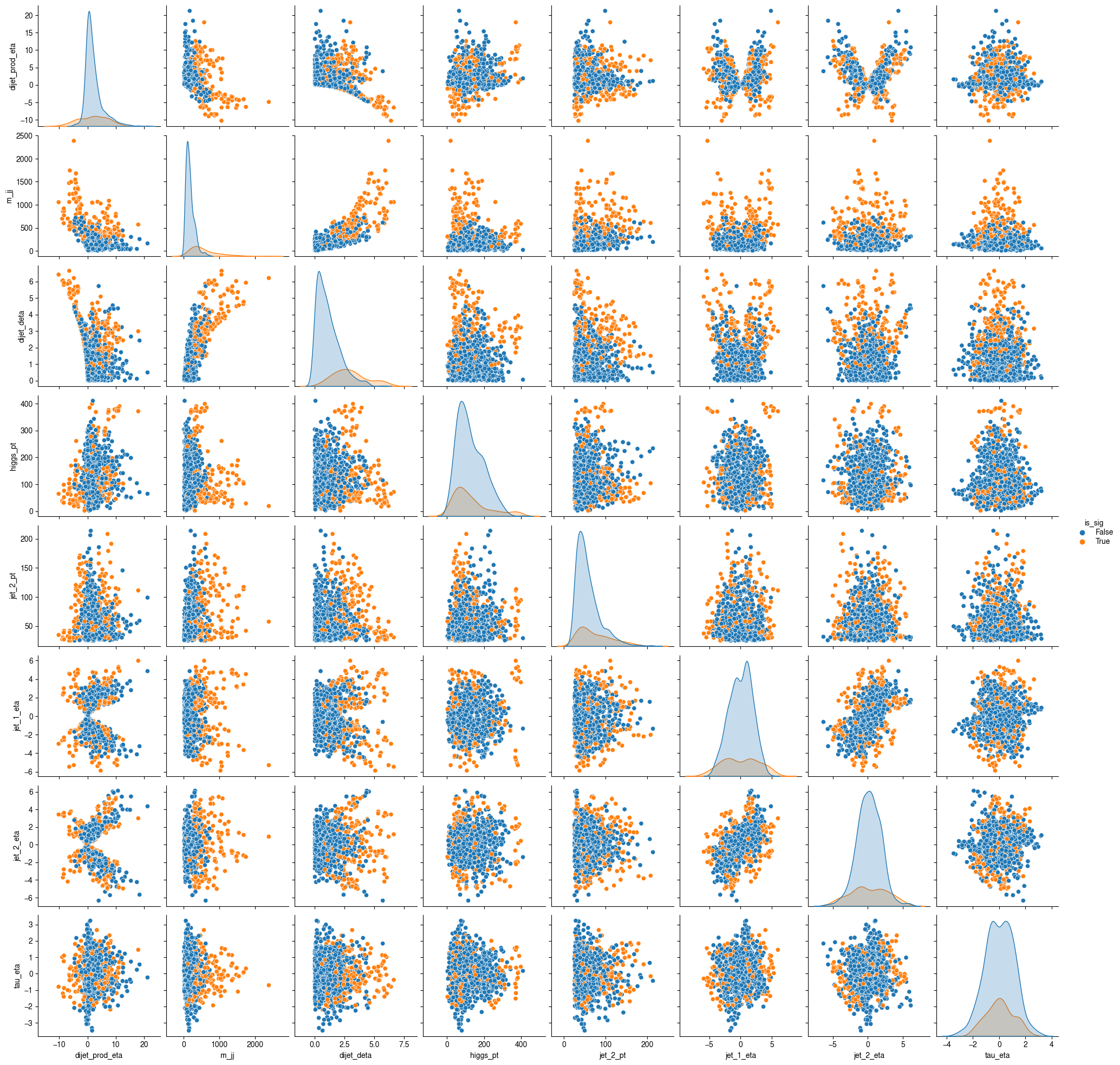
sample_df = df.sample(frac=1000 / len(df)).compute()
sns.pairplot(sample_df, vars=input_var, hue="is_sig")
None
[11]:
def model():
m = Sequential()
m.add(Dense(units=15, activation='relu', input_dim=len(input_var)))
m.add(Dense(units=5, activation='relu'))
m.add(Dense(units=2, activation='softmax'))
m.compile(loss='categorical_crossentropy',
optimizer=SGD(lr=0.1),
weighted_metrics=['categorical_accuracy'])
return m
cv = ClassicalCV(5, frac_var='random')
net = HepNet(model, cv, EstimatorNormalizer, input_var, output_var)
[12]:
sig_wf = len(p_sig.selection(df).weight) / p_sig.selection(df).weight.sum()
ztt_wf = len(p_ztt.selection(df).weight) / p_ztt.selection(df).weight.sum()
[13]:
net.fit(df.compute(), epochs=150, verbose=0, batch_size=2048,
weight=Variable("weight", lambda d: d.weight * (d.is_sig * sig_wf + d.is_ztt * ztt_wf)))
2023-08-02 16:28:57.394711: W tensorflow/compiler/xla/stream_executor/platform/default/dso_loader.cc:64] Could not load dynamic library 'libcuda.so.1'; dlerror: libcuda.so.1: cannot open shared object file: No such file or directory
2023-08-02 16:28:57.394754: W tensorflow/compiler/xla/stream_executor/cuda/cuda_driver.cc:265] failed call to cuInit: UNKNOWN ERROR (303)
2023-08-02 16:28:57.394779: I tensorflow/compiler/xla/stream_executor/cuda/cuda_diagnostics.cc:156] kernel driver does not appear to be running on this host (build-21485799-project-648611-freeforestml): /proc/driver/nvidia/version does not exist
2023-08-02 16:28:57.395010: I tensorflow/core/platform/cpu_feature_guard.cc:193] This TensorFlow binary is optimized with oneAPI Deep Neural Network Library (oneDNN) to use the following CPU instructions in performance-critical operations: AVX2 AVX512F FMA
To enable them in other operations, rebuild TensorFlow with the appropriate compiler flags.
WARNING:absl:`lr` is deprecated, please use `learning_rate` instead, or use the legacy optimizer, e.g.,tf.keras.optimizers.legacy.SGD.
WARNING:absl:`lr` is deprecated, please use `learning_rate` instead, or use the legacy optimizer, e.g.,tf.keras.optimizers.legacy.SGD.
WARNING:absl:`lr` is deprecated, please use `learning_rate` instead, or use the legacy optimizer, e.g.,tf.keras.optimizers.legacy.SGD.
WARNING:absl:`lr` is deprecated, please use `learning_rate` instead, or use the legacy optimizer, e.g.,tf.keras.optimizers.legacy.SGD.
WARNING:absl:`lr` is deprecated, please use `learning_rate` instead, or use the legacy optimizer, e.g.,tf.keras.optimizers.legacy.SGD.
[14]:
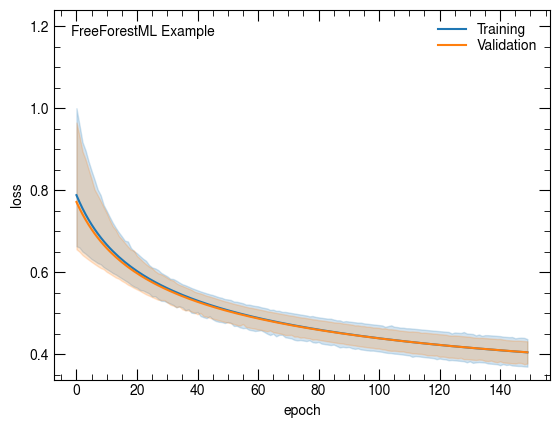
sns.lineplot(x='epoch', y='loss', data=net.history, label="Training")
sns.lineplot(x='epoch', y='val_loss', data=net.history, label="Validation")
plt.ylabel("loss")
atlasify(False, "FreeForestML Example")
None
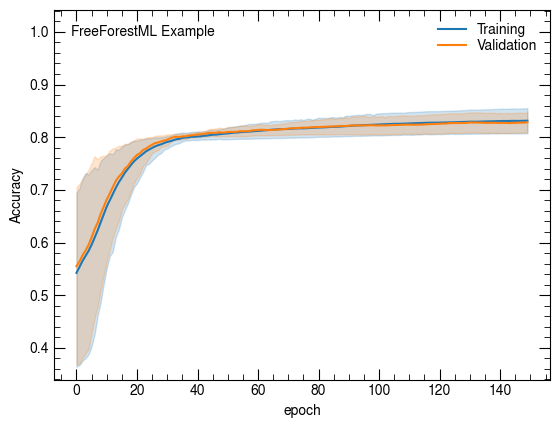
Accuracy
[15]:
sns.lineplot(x='epoch', y='categorical_accuracy', data=net.history, label="Training")
sns.lineplot(x='epoch', y='val_categorical_accuracy', data=net.history, label="Validation")
plt.ylabel("Accuracy")
atlasify(False, "FreeForestML Example")
None
[16]:
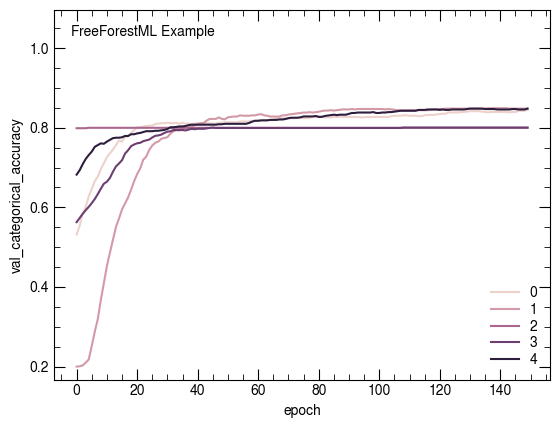
sns.lineplot(x='epoch', y='val_categorical_accuracy', data=net.history, hue="fold")
plt.legend(loc=4)
atlasify(False, "FreeForestML Example")
None
[17]:
out = net.predict(df.compute(), cv='test')
out['pred_sig'] = out.pred_is_sig >= 0.5
162/162 [==============================] - 0s 1ms/step
162/162 [==============================] - 0s 1ms/step
162/162 [==============================] - 0s 1ms/step
162/162 [==============================] - 0s 1ms/step
162/162 [==============================] - 0s 1ms/step
[18]:
c_pred_sig = Process("Signal", lambda d: d.pred_is_sig >= 0.5)
c_pred_ztt = Process(r"$Z\rightarrow\tau\tau$", lambda d: d.pred_is_sig < 0.5)
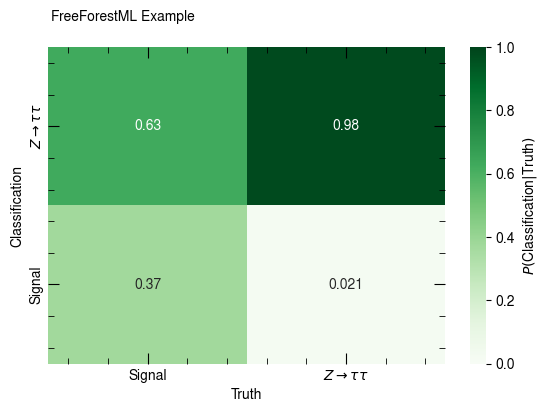
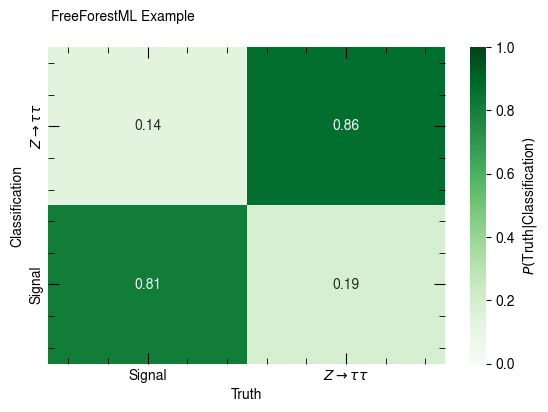
confusion_matrix(out, [p_sig, p_ztt], [c_pred_sig, c_pred_ztt],
x_label="Truth", y_label="Classification", annot=True, weight="weight")
confusion_matrix(out, [p_sig, p_ztt], [c_pred_sig, c_pred_ztt], normalize_rows=True,
x_label="Truth", y_label="Classification", annot=True, weight="weight")
None
Export to lwtnn
In order to use the network in lwtnn, we need to export the neural network with the export() method. This export one network per fold. It is the reposibility of the use to implement the cross validation in the analysis framework.
[19]:
net.export("lwtnn")
[20]:
!ls lwtnn*
lwtnn.sh lwtnn_arch_3.json lwtnn_vars_2.json lwtnn_wght_1.h5
lwtnn_arch_0.json lwtnn_arch_4.json lwtnn_vars_3.json lwtnn_wght_2.h5
lwtnn_arch_1.json lwtnn_vars_0.json lwtnn_vars_4.json lwtnn_wght_3.h5
lwtnn_arch_2.json lwtnn_vars_1.json lwtnn_wght_0.h5 lwtnn_wght_4.h5
The final, manuel step is to run the lwtnn’s converter using the shortcut script test.sh.
[ ]: